Chong Li
2007.9-2011.5 Yantai University, Pharmacy, B.S.
2012.8-2018.5 University of Miami, Miller School of Medicine (U.S.), Human genetics and Genomics, Ph.D.
2018.8-2024.8 IMBA-Institute of Molecular Biotechnology of the Austrian Academy of Sciences, Postdoc (Jürgen Knoblich Group)
2024.8-Current Chinese Institute for Brain Research, Beijing (CIBR), Assistant Investigator
Our laboratory is dedicated to unraveling the complexities of the human brain, an organ distinguished by its 100 billion neurons, over 3,000 cell types, and approximately 100 trillion synaptic connections. This remarkable complexity makes the human brain the most sophisticated among all primates. Our research seeks to answer pivotal questions about the unique developmental principles of the human brain, the precise regulatory mechanisms of neural stem cell fate, and the implications of these processes for brain disorders such as autism.
Core Research Areas:
1. Fundamental Principals of Human Brain Development: We investigate how human brain development differs from that of other species and how these differences underpin its complex structure and functionalities. Our studies focus on identifying the genetic factors that influence brain architecture and operational dynamics.
2. Neural Stem Cell Regulation: Understanding the mechanisms that dictate neural stem cell behavior is crucial for deciphering brain development. Our work examines how these cells differentiate, proliferate, contribute to, and maintain the diversity of cell types found in the brain, providing insights into both normal development and pathological conditions.
3. Brain Disorder Mechanisms: By studying deviations in normal brain development, we aim to identify the root causes of brain disorders. Our research involves characterizing the molecular and cellular changes that lead to conditions like autism and cognitive deficits, offering potential pathways for therapeutic intervention.
Methodological Approaches:
1. Human Brain Organoids: Utilizing brain organoids derived from human pluripotent stem cells (PSCs), we create models that mimic the development and organization of the human brain in vitro. This approach allows us to study the developing processes of the human brain in real time (Figure 1).
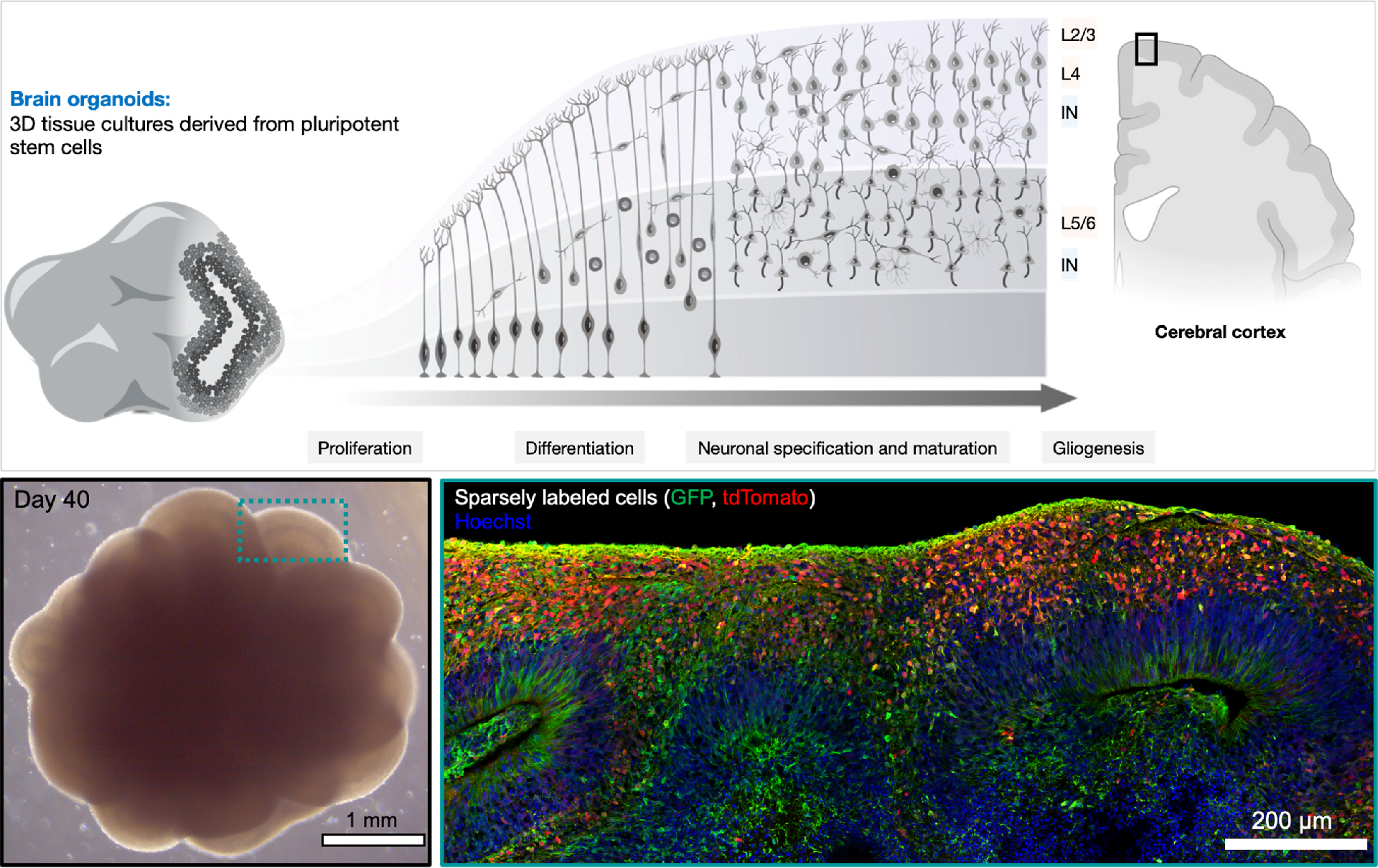
Figure 1
2. CRISPR Gene Editing: We employ CRISPR technology to edit the genomes in multiple ways, enabling us to investigate the effects of specific genetic modifications on brain development and function (Figure 2).
3. High-Throughput Screening: Our lab uses high-throughput techniques to systematically analyze and compare large number of genes in parallel, accelerating the discovery of factors that influence brain development and disease (Figure 2).
4. Single-Cell Genomics: By analyzing individual cells through single cell genomics technology (e.g., transcriptome and epigenome), we gain a deeper understanding of the cellular and molecular diversity within the brain and how it changes in disease states (Figure 2).
5. Quantitative Biology: We integrate quantitative methods to measure and model the dynamics of brain development, providing a comprehensive view of the processes that shape the human brain (Figure 2).
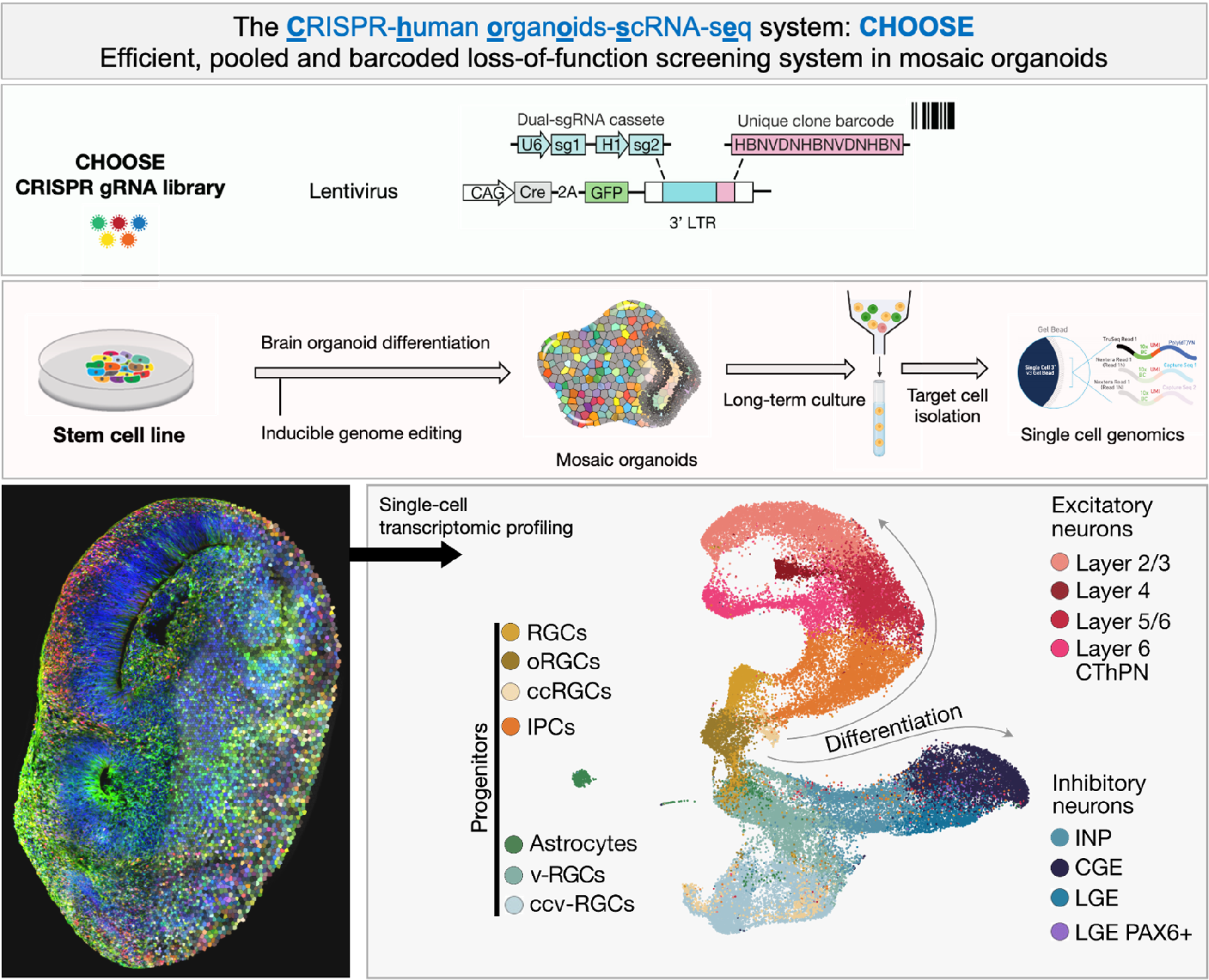
Figure 2
Our lab is committed to addressing some of the most urgent challenges in brain research. Through a combination of innovative model systems and advanced technological approaches, we explore the fundamental principles that govern brain development and contribute to disease processes, aiming to translate our findings into therapeutic strategies. We are also committed to creating an innovative and diverse research environment, encouraging lab members to fully explore their scientific interests. We warmly welcome passionate PhD students, postdocs, interns, and research assistants to join us on an enjoyable journey into brain research.
04/2023 Best Talk Award, Gordon Research Conference, Functional Genomics of Human Brain Development and Disease
11/2017 Best Research Award, Medical Faculty Association, University of Miami
09/2017 American Society for Cell Biology Travel Award
11/2016 Dr. Louis J. Elsas Award in Biochemical Genetics, Dr. John T. Macdonald Foundation
10/2016 Medical Faculty Association Travel Award, The Margaret Whelan Fund
02/2016 Snyder-Robinson Foundation Predoctoral Fellowship, The Snyder-Robinson Foundation
12/2015 Lois Pope LIFE Fellowship Award, Lois Pope LIFE Foundation
1. Li CV#, Knoblich JA#. Advancing autism research: Insights from brain organoid modeling. Curr Opin Neurobiol. 2025 Apr 24;92:103030. doi: 10.1016/j.conb.2025.103030(# co-corresponding)
2. Bosone C*, Castaldi D*, Burkard T R, Guzman S J, Wyatt T, Cheroni C, Caporale N, Bajaj S, Bagley J A, Li C, Sorre B, Villa E, Testa G, Krenn V, and Knoblich J A. (2024) A polarized FGF8 source specifies frontotemporal signatures in spatially oriented cell populations of cortical assembloids Nature Methods 21, 2147–2159
3. Li C*#, Fleck J S*, Martins-Costa C, Burkard T R, Stuempflen M, Vertesy Á, Peer A M, Esk C, Elling U, Kasprian G, Corsini N S, Treutlein B# and Knoblich J A#. (2023) Single-cell brain organoid screening identifies developmental defects in autism. Nature 621, 373–380 (*equal contribution, # co-corresponding)
4. Liu J, Tao X, Zhu Y, Li C, Ruan K, Diaz-Perez Z, Rai P, Wang H, Zhai RG. (2021) NMNAT promotes glioma growth through regulating post-translational modifications of P53 to inhibit apoptosis. Elife. 2021 Dec 17;10:e70046. doi: 10.7554/eLife.70046.
5. Ma X, Zhu Y, Lu J, Xie J, Li C, Shin WS, Qiang J, Liu J, Dou S, Xiao Y, Wang C, Jia C, Long H, Yang J, Fang Y, Jiang L, Zhang Y, Zhang S, Zhai RG, Liu C, Li D. (2020) Nicotinamide mononucleotide adenylyltransferase uses its NAD+ substrate-binding site to chaperone phosphorylated Tau. Elife. Apr 6;9:e51859. doi: 10.7554/eLife.51859.
6. Zhu Y, Li C, Tao X, Brazill JM, Park J, Diaz-Perez Z, Zhai RG (2019) Nmnat restores neuronal integrity by neutralizing mutant Huntingtin aggregate-induced progressive toxicity. PNAS, 116 (38), 19165-19175
7. Park J, Zhu Y, Tao X, Brazill JM, Li C, Wuchty S, Zhai RG (2019) Microrna mir-1002 enhances Nmnat-mediated stress response by modulating alternative splicing. iScience, 19:1048-1064.
8. Li C*, Bademci G*, Subasioglu A, Diaz-Horta O, Zhu Y, Liu J, Mitchell TG, Abad C, Seyhan S, Duman D, Cengiz FB, Tokgoz-Yilmaz S, Blanton SH, Farooq A, Walz K, Zhai RG, Tekin M (2019) Dysfunction of GRAP, encoding the GRB2-related adaptor protein, is linked to sensorineural hearing loss. (*equal contribution), PNAS, 116: 1347-1352
9. Brazill J, Zhu Y, Li C, Zhai RG (2018) Quantitative cell biology of neurodegeneration in Drosophila through unbiased analysis of fluorescently tagged proteins using ImageJ. J. Vis. Exp. (138), e58041, doi:10.3791/58041.
10. Li C, Brazill J, Liu S, Bello C, Zhu Y, Morimoto M, Cascio L, Pauly R, Diaz-Perez Z, Malicdan MCV, Wang H, Boccuto L, Schwartz CE, Gahl WA, Boerkoel CF, Zhai RG (2017) Spermine synthase deficiency causes lysosomal dysfunction and oxidative stress in models of Snyder-Robinson syndrome. Nature Communications, 8(1):1257
11. Gall T, Valkanas E, Bello C, Markello T, Adams C, Bone WP, Brandt AJ, Brazill JM, Carmichael L, Davids M, Davis J, Diaz-Perez Z, Draper D, Elson J, Flynn ED, Godfrey R, Groden C, Hsieh C-K, Fischer R, Golas GA, Guzman J, Huang Y, Kane MS, Lee E, Li C, Links AE, Maduro V, Malicdan MCV, Malik FS, Nehrebecky M, Park J, Pemberton P, Schaffer K, Simeonov D, Sincan M, Smedley D, Valivullah Z, Wahl C, Washington N, Wolfe LA, Xu K, Zhu Y, Gahl WA, Tifft CJ, Toro C, Adams DR, He M, Robinson PN, Haendel MA, Zhai RG and Boerkoel CF (2017) Defining disease, diagnosis, and translational medicine within a homeostatic perturbation paradigm: The NIH Undiagnosed Diseases Program experience. Front. Med., 4:62.
12. Brazill JM*, Li C*, Zhu Y*, Zhai RG (2017) NMNAT: It’s an NAD+ Synthase… It’s a chaperone… It’s a neuroprotector. Curr Opin Genetics Dev., 44:156–162. (*Equal contribution).
13. Ruetenik A, Ocampo A, Ruan K, Zhu Y, Li C, Zhai RG and Barrientos A (2016) Attenuation of polyglutamine-induced toxicity by enhancement of mitochondrial OXPHOS in yeast and fly models of aging. Microbial Cell 3(8):338-351.
14. Ruan K, Zhu Y*, Li C*, Brazill J, and Zhai RG (2015) Alternative splicing of Drosophila Nmnat acts as a switch to enhance neuroprotection under stress. Nature Communications 6:10057 (*Equal contribution)
15. Li C*, Wang T*, Jiang N, Yu PF, Du Y, Ren RT, and Fu FH (2012) Steady and fluctuant methods of inhibition of acetylcholinesterase differentially regulate neurotrophic factors in the hippocampus of juvenile mice. Exp Ther Med, 3(2):269-272. (*Equal contribution)



